Research
Our research program advances precision medicine through a data-centric approach, developing and applying computational methods to analyze diverse biomedical datasets across multiple scales — from single cells to patient populations.
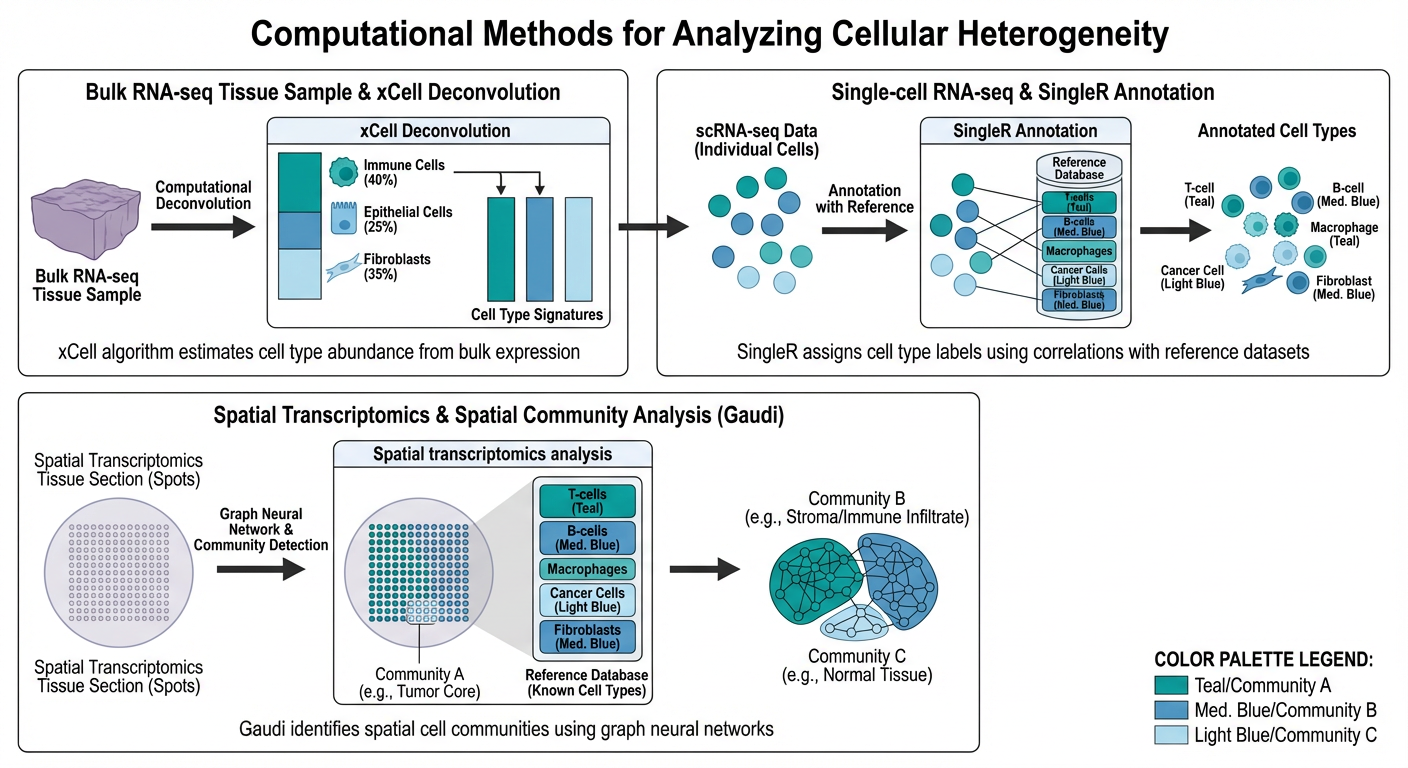
Computational Methods for Cellular Heterogeneity
Cellular heterogeneity in tissues represents one of the fundamental challenges in modern biomedical research. Our lab develops computational methods that bridge different data modalities and scales to characterize this heterogeneity. In bulk RNA-seq, we built upon xCell to develop xCell 2.0, a flexible deconvolution framework that consistently outperforms other methods. For single-cell analysis, we developed a hierarchical deep-phenotyping framework building on SingleR, and CellMentor for cell-type aware dimensionality reduction. Our spatial transcriptomics work with the Gaudi framework identifies cellular communities using graph neural networks. We also developed SLAYER for synthetic lethal interactions and Fine-Pruning for making AI models more accessible in clinical settings.
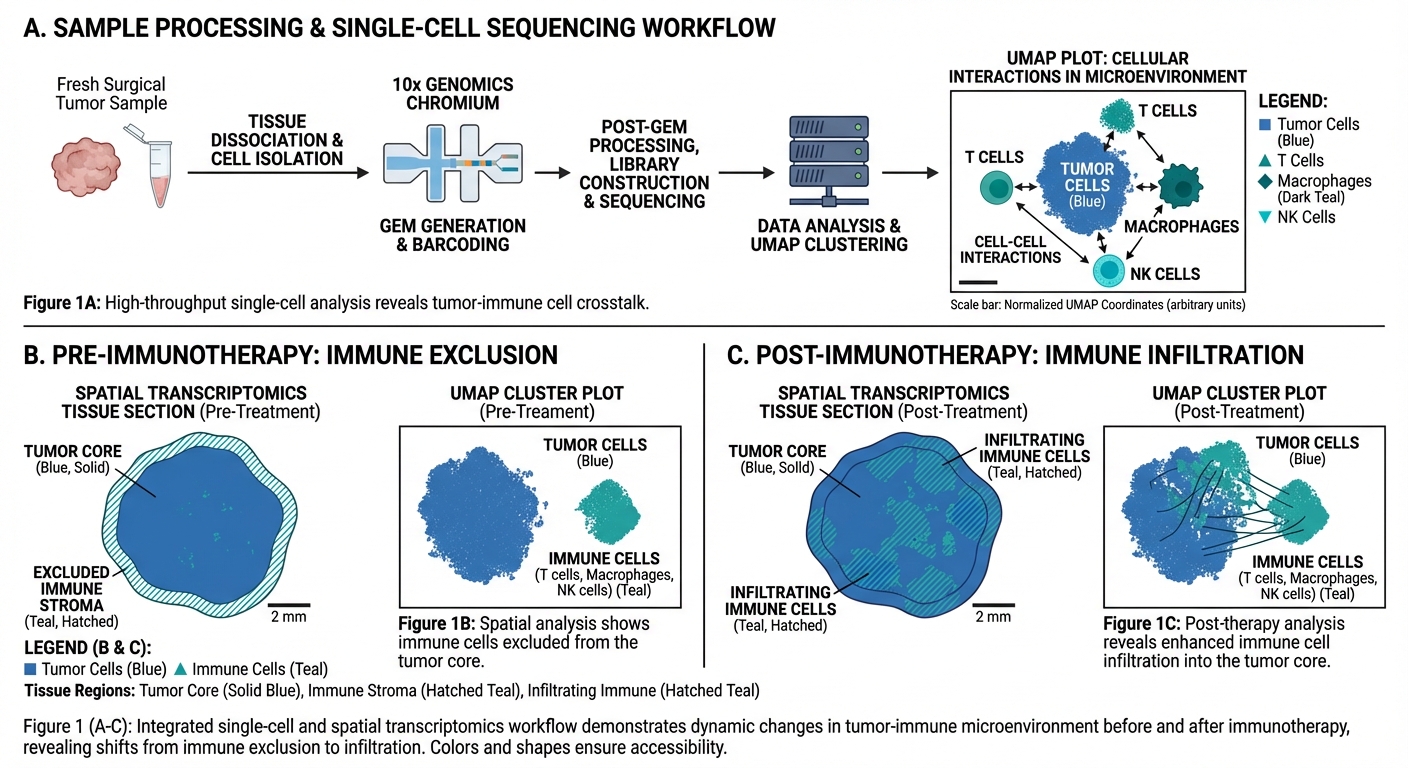
Cellular Dynamics in Health & Disease
Beyond method development, we apply computational approaches to understand how cells behave and interact in various biological contexts. We established protocols for collecting and analyzing fresh surgical samples from Rambam Medical Center, processing over 50 samples through scRNA-seq and spatial transcriptomics. A key focus is our comprehensive analysis of tumor microenvironment changes during immunotherapy across 326 samples from 163 patients. We also investigate brain metastases after immunotherapy, pre-metastatic niches in head and neck cancer, age-dependent neuroblastoma prognosis, and neuroimmunological responses in myasthenia gravis.
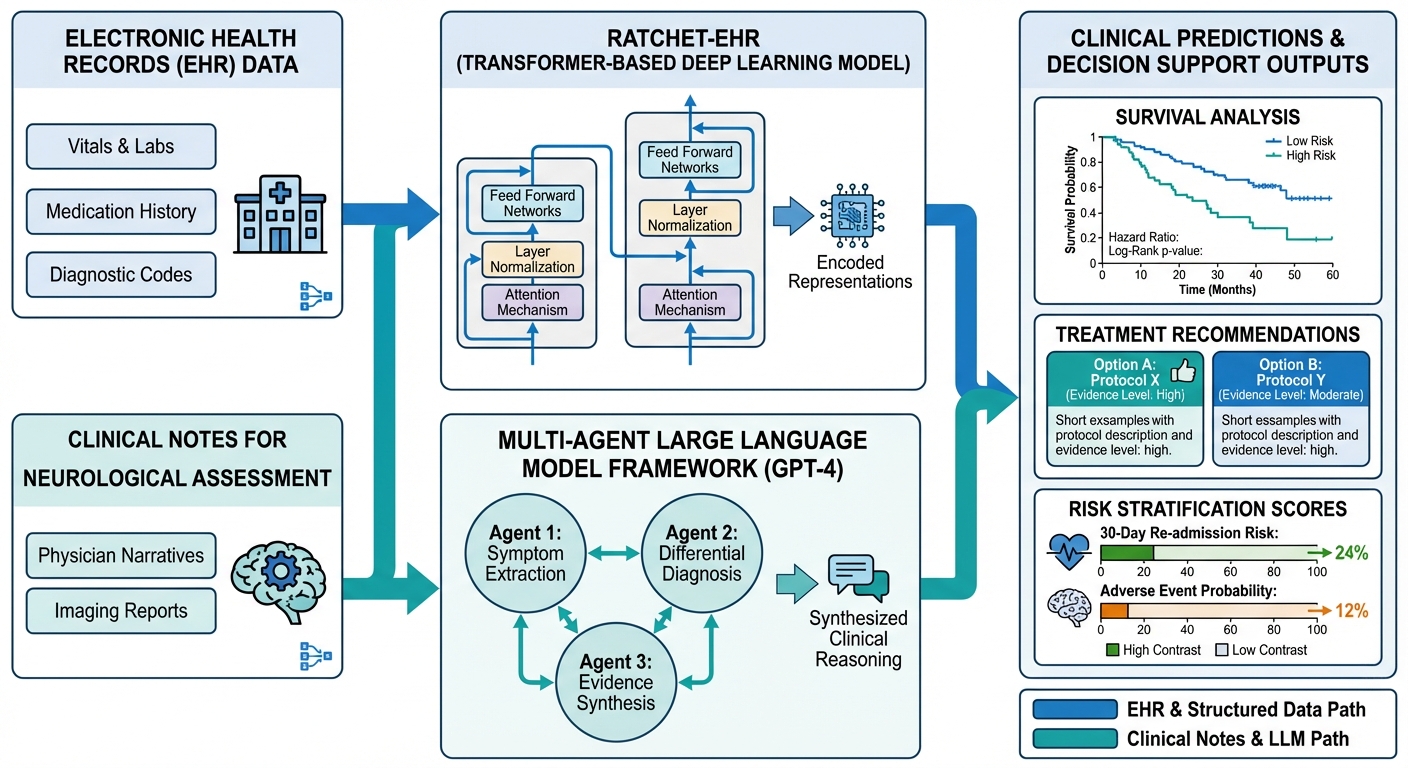
Clinical Decision-Making through AI
The third pillar of our research translates computational insights into clinical applications. Our deep learning approaches for electronic health records include RatchetEHR for ICU data analysis and STRAFE for survival prediction. We evaluate large language models for clinical decision support, including GPT-4 for ischemic stroke management and multi-agent frameworks for neurological reasoning. Our real-world evidence studies have yielded practice-changing insights in metastatic pancreatic cancer (JAMA Network Open) and melanoma immunotherapy (BJC Reports). We also develop predictive models for nursing workforce optimization, nutritional risk assessment, and cancer screening.